OnRamp’s ROSALIND and Active Motif offer an improved ChIP-Seq data analysis experience for researchers with state-of-the-art dynamic track plots and advanced collaboration capabilities.
FOR IMMEDIATE RELEASE
San Diego, California – December 12, 2018 - OnRamp BioInformatics, a genomics company focused on scientist-friendly bioinformatics software solutions, today announced they are working with Active Motif to improve accuracies in data analysis for ChIP-Seq experiments.
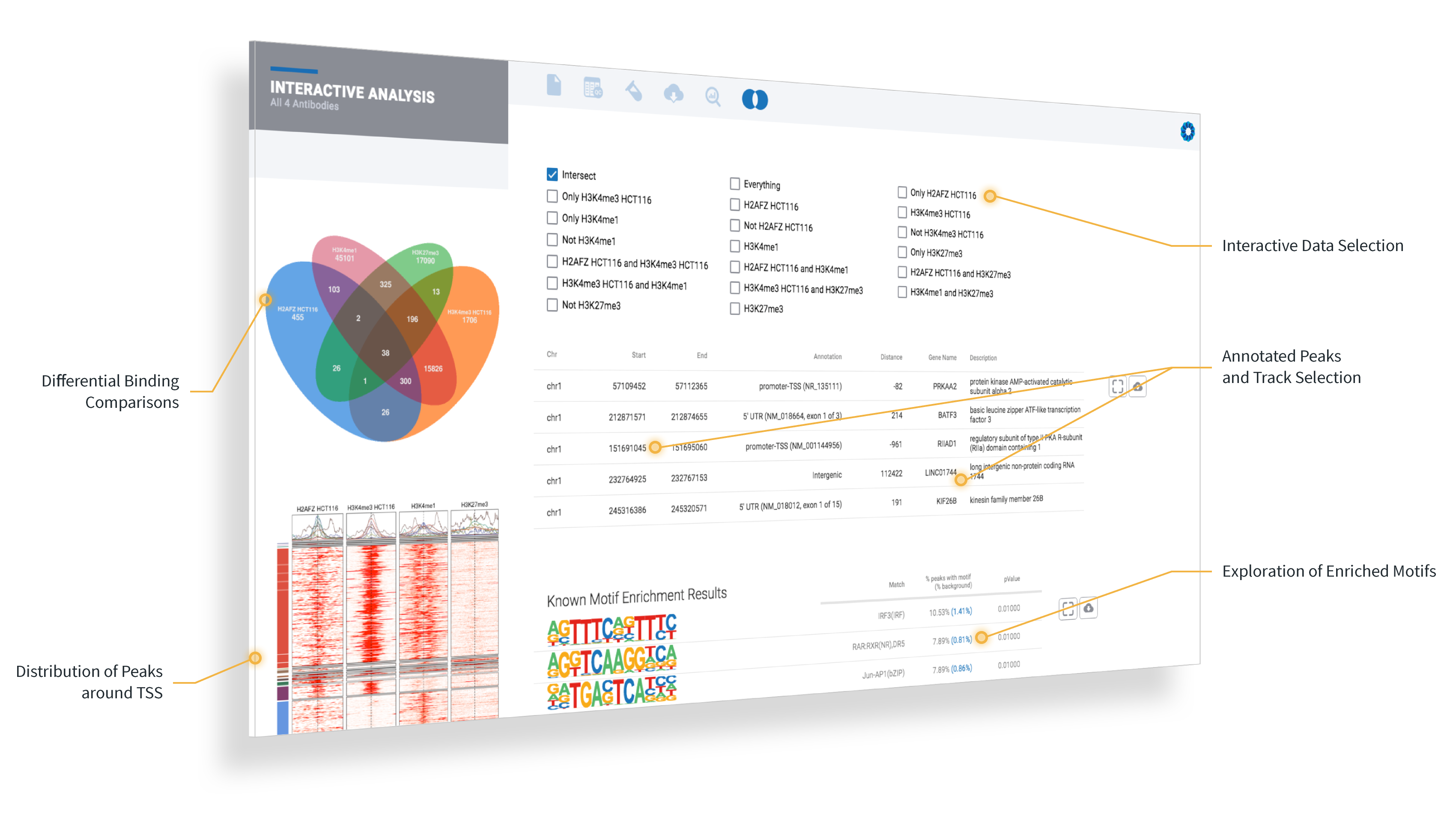
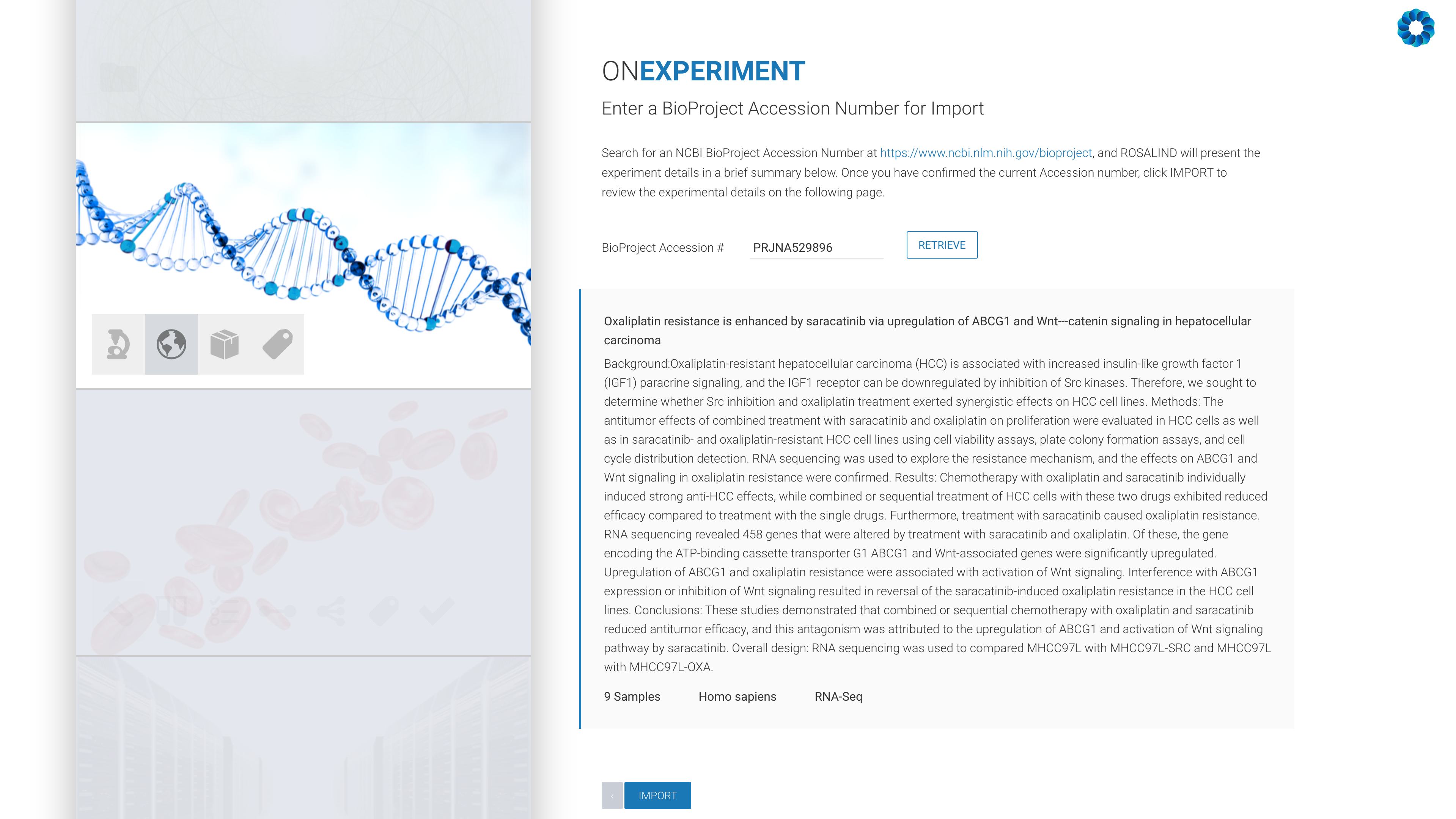
OnRamp’s flagship software, ROSALIND, provides a point-and-click experience to speed up and simplify the process of genomic data analysis from experiment setup to QC and interactive data visualization to interpretation. With this integration, researchers will now benefit from an optimized analysis that includes the unique characteristics of Active Motif kits and antibodies, including Unique Molecular Identifiers (UMI) and spike-in solutions.
Tweet: Combining the simplicity of Active Motif kits with the ROSALIND online analysis platform allows even beginners to complete an entire ChIP-Seq experiment with ease

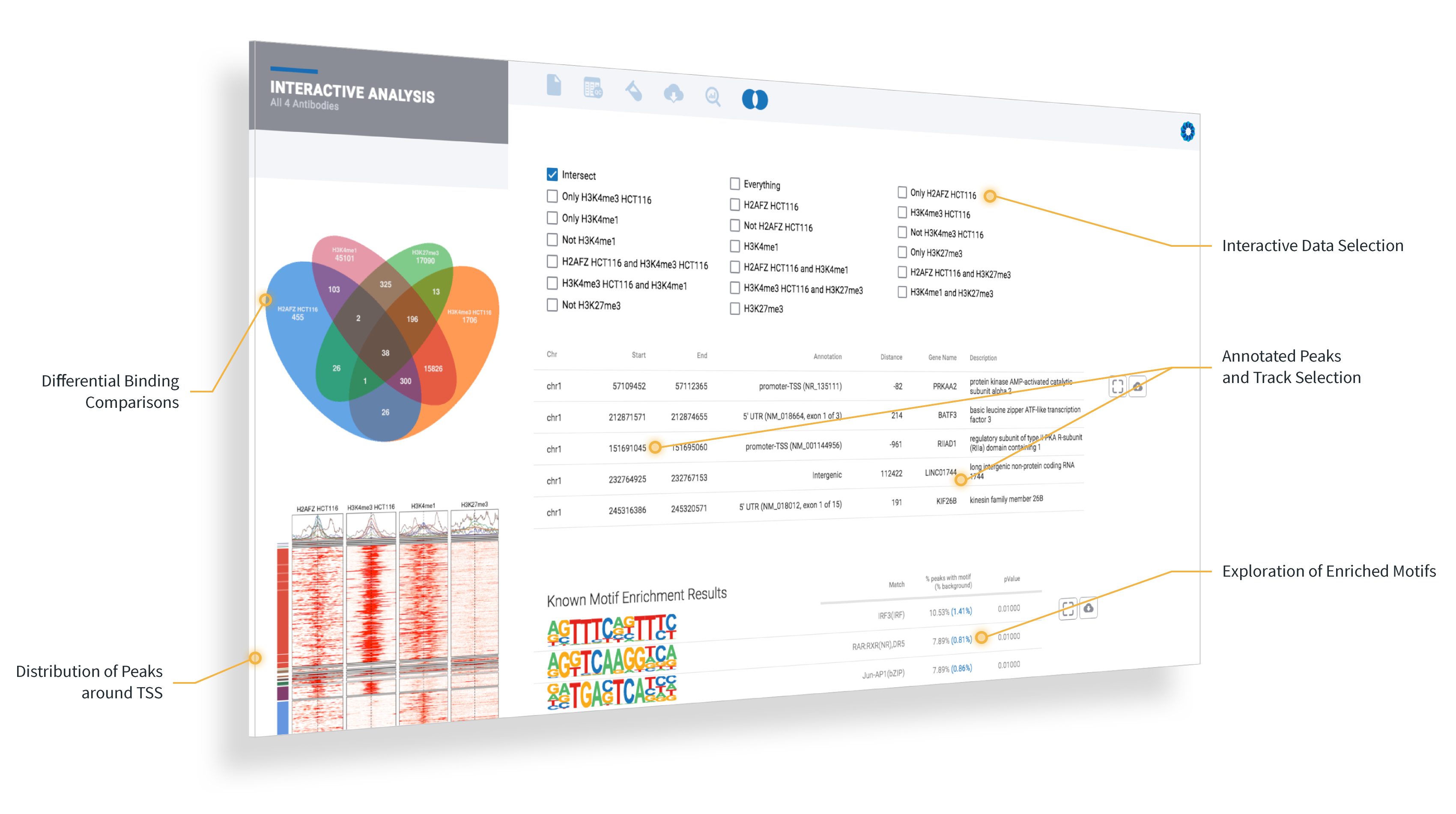
Active Motif customers will now be able to rapidly interpret their data with ROSALIND’s identification of the significant peaks and known or de novo binding motifs, as well as a functional enrichment analysis based on “close-by genes” using over 20 knowledgebases like GO, WikiPathways, MSigDB, Pfam, Interpro, and more.
Scientists and researchers can then share their experiment using a ROSALIND collaborative space to virtually work alongside colleagues and explore the results without having to exchange their files.
“Combining the simplicity of our kits with a straightforward online analysis solution allows even beginners to complete an entire ChIP-Seq experiment with ease,” says Ted DeFrank, CEO of Active Motif. “It is our hope that offering a way to analyze ChIP-Seq data with speed and improved accuracy will enable users to publish new findings much more quickly.”
“Accelerating discoveries in Epigenetics is critical for drug discovery and disease research,” says Tim Wesselman, CEO of OnRamp BioInformatics. “We’re excited to partner with Active Motif to empower scientists who use their high-quality kits and antibodies with an intuitive ROSALIND analysis, so they can focus on their insights and discoveries, while saving time and removing the frustration that comes with trying to generate them.”
About OnRamp BioInformatics
Based in San Diego, OnRamp.Bio is a genomics company providing bioinformatics analysis services and cost-effective platforms for the computing and storage of genomic datasets. OnRamp.Bio empowers researchers, biologists and scientists with genomic answers and insights that they can understand and put to action. To learn more, check out our website www.onramp.bio, find us on Twitter (@OnRampBio) and Facebook, or visit our blog.
About Active Motif
Active Motif is the industry leader in developing and delivering innovative tools to enable epigenetics and gene regulation research for the life science, clinical and pharmaceutical and drug discovery communities. The company has a world-leading portfolio of assays, genome-wide services and validated antibodies for use in ChIP-Seq. Active Motif operates globally through its corporate headquarters in Carlsbad, California, regional headquarters in Belgium, Japan and China, as well as a worldwide network of sales and support offices. Active Motif applies a multi-disciplinary approach to create new and modify existing technologies to meet the current and future needs of life science researchers. To learn more please visit www.activemotif.com.
Media Inquiries
+1 855-766-7267 ext. 708