This new web-based system, developed with NIH funding, will aid in monitoring emerging SARS-CoV-2 variants and the performance of diagnostic tests
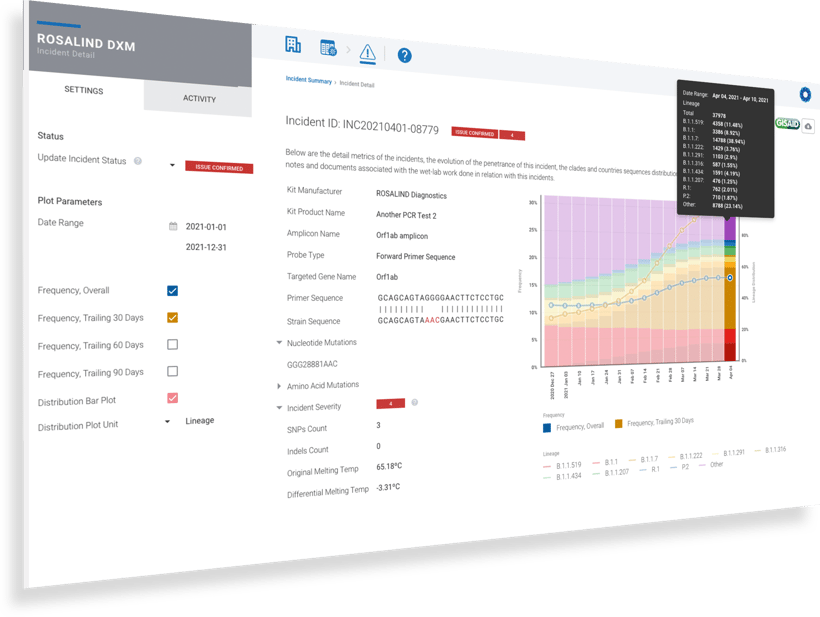
SAN DIEGO, CA – July 12, 2021 – ROSALIND®, a software leader in extracting meaningful insights from diverse pools of life science data, today announced the launch of a web-based platform for monitoring the emergence of SARS-CoV-2 variants and evaluating the effects on the performance of diagnostic tests. The new platform, called the ROSALIND Diagnostic Monitoring (DxM) system, is available starting today and will be used by diagnostic test regulators to monitor assay performance against the rapidly evolving SARS-CoV-2 variants.
With the rapid emergence of SARS-CoV-2 variants, global awareness has led to rising concern over the detection of new viral strains using the existing diagnostic assays. The U.S. Food and Drug Administration (FDA) issued an alert in January that false-negative results may occur with any molecular test for the detection of SARS-CoV-2 if a mutation occurs in the part of the virus’ genome assessed by that test. The agency, urged test makers to perform periodic in silico assessment (computer simulations) of molecular tests to identify impacts on performance.
ROSALIND developed the DxM system with support from the Rapid Acceleration of Diagnostics (RADxSM) Tech program, part of the National Institutes of Health’s RADx initiative launched in April 2020 to speed innovation in the development, commercialization, and implementation of technologies for COVID-19 testing. Recognizing the importance of a centralized bioinformatics system, RADx Tech issued a request for proposals to develop and implement a bioinformatics pipeline with a user-friendly web-based interface that would provide regular updates of the frequency of the variants, mapping/alignment of the probe/detection sequences of the NIH RADx projects and to assess the effect of the variants on the performance of the tests. ROSALIND was awarded the contract in late February 2021 and today released the first version of the new platform.
“We are honored to be a collaborator in this important project and inspired to support the NIH in the fight against COVID-19. The DxM system has the potential to play a crucial role in identifying risks to diagnostic tests from this evolving virus, and aids test manufacturers and agencies with consistent analyses and recommendations,” said Tim Wesselman, CEO of ROSALIND. “This collaborative project is an extension of our vision to empower scientists to extract meaningful insights from genomic data. Leveraging our existing ROSALIND platform has enabled us to respond with incredible speed while providing a highly secure and consistent analysis system that will be utilized across many organizations and agencies.”
ROSALIND is working with SARS-CoV-2 diagnostic manufacturers to provide system access and provide ongoing analysis and reporting based on sequencing data from scientists and laboratories across the globe.
“With all the complexity in the SARS-CoV-2 sequence data and the many different types of diagnostic tests — PCR, antigen, CRISPR, and more — we understood right from the start that diagnostic manufacturers and agencies needed an intuitive platform where they could define their tests and immediately detect potential issues against all available sequence data,” said Jean Lozach, Chief Science Officer of ROSALIND. “ROSALIND continuously queries each test against a database of all available sequencing data and emerging variants. When a risk is detected, an alert is sent via a secure message with a report out of the findings. After receiving the alert, users can engage with interactive dashboards to investigate variant details and lookup specimens for wet-lab verification of in silico assessments.”
The RADx initiative has supported 150 companies in advancing their diagnostic technologies and the development of 16 COVID-19 tests that have received FDA authorization.
This project has been funded in whole or in part through a $1.5 billion investment of federal funds from the National Institute of Biomedical Imaging and Bioengineering, National Institutes of Health, and the Department of Health and Human Services. ROSALIND is supporting this project as a subcontractor of VentureWell under Contract 75N92020P00171.
For more information on the DxM System and other RADx funded programs, visit https://www.rosalind.bio/dxm, or
https://www.nih.gov/research-training/medical-research-initiatives/radx.
About ROSALIND
Based in San Diego, ROSALIND enables the world’s life science researchers to derive more value from their data and complete projects faster. We believe the potential of biomedical and multi-omic data is maximized when it is accessible, interactive, and shareable. Extracting meaningful insights from pools of data takes a truly interoperable ecosystem that breaks down silos of complex information and sparks collaboration. The ROSALIND platform harnesses the power of scientific collaboration, cloud computing, and data science to enable integrated analyses that unify knowledge through an intuitive user interface that is built for any researcher. For more information, please visit www.rosalind.bio.
Related Posts
-
For all Research affected by Social Distancing: Remote Science in the Era of Social Distancing demonstrates how working remotely does not mean that science must stop.
-
For Scientists on ROSALIND: Collaborating on Genomic Data is a walkthrough on how to set up real-time collaboration on genomic data (like Google Docs). Read the post here
-
For Directors and Managers: The Value is in Collaboration (not the pipeline) discusses how collaboration is key to empowering research in your lab.
-
For every Researcher, Public Data exploration shows how we have made it 100x easier to work with public data than ever before.
- To learn more about our latest capabilities, Empowering Scientists with the ROSALIND Platform is an overview of all analysis types, species and features currently available to the general public on ROSALIND.









.png)