Our latest ROSALIND update v3.16 is now live and delivers enhancements for collaboration, 10X Single Cell RNA (scRNA-seq), Pathway Enrichment and more.
Recent posts by Jeremy Davis-Turak
4 min read
ROSALIND v3.16: Enhanced Collaboration and Pathways
By Jeremy Davis-Turak on Jun 26, 2020 7:28:36 PM
Topics: BioInformatics Rosalind Public Data Collaboration COVID-19 Enterprise
3 min read
Active Motif & Rosalind Announce Partnership
By Jeremy Davis-Turak on Dec 12, 2018 7:37:00 AM
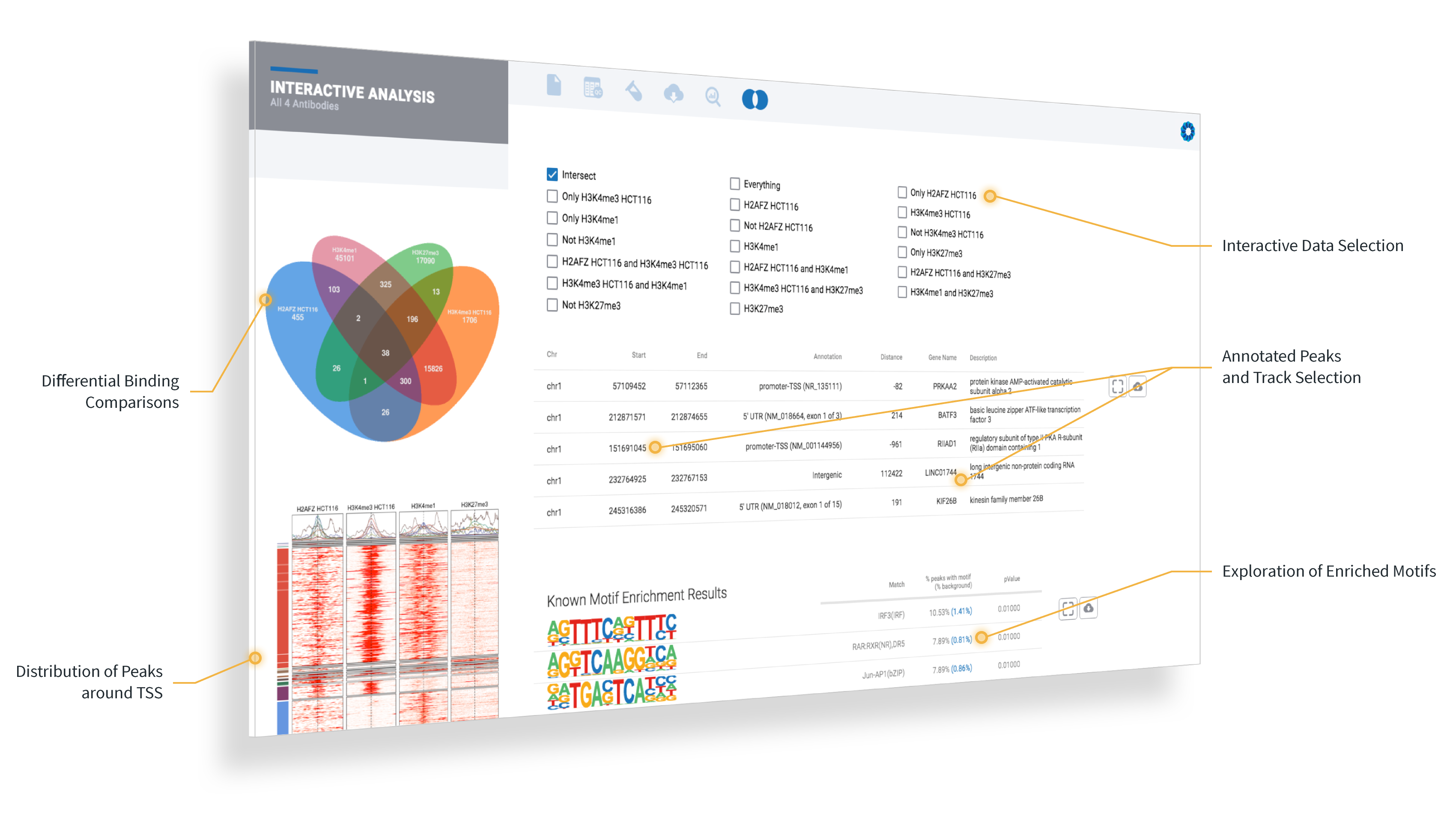
OnRamp’s ROSALIND and Active Motif offer an improved ChIP-Seq data analysis experience for researchers with state-of-the-art dynamic track plots and advanced collaboration capabilities.
FOR IMMEDIATE RELEASE
San Diego, California – December 12, 2018 - OnRamp BioInformatics, a genomics company focused on scientist-friendly bioinformatics software solutions, today announced they are working with Active Motif to improve accuracies in data analysis for ChIP-Seq experiments.
OnRamp’s flagship software, ROSALIND, provides a point-and-click experience to speed up and simplify the process of genomic data analysis from experiment setup to QC and interactive data visualization to interpretation. With this integration, researchers will now benefit from an optimized analysis that includes the unique characteristics of Active Motif kits and antibodies, including Unique Molecular Identifiers (UMI) and spike-in solutions.
Topics: Big Data Personalized Medicine Cancer BioInformatics Biology Rosalind advaita
1 min read
View Our Meta Analysis Webinar
By Jeremy Davis-Turak on May 9, 2018 9:19:00 AM
Building a "Comparison of Comparisons" Across Multiple Experiments using Meta-Analysis on ROSALIND
Topics: Transcriptome BioInformatics Webinar RNAseq
4 min read
Decoding Human Health
By Jeremy Davis-Turak on May 3, 2018 9:10:20 AM
OnRamp and Advaita announce a Strategic Partnership to bring collaboration and widespread accessibility to genomic data analysis
San Diego, CA – May 3, 2018 – OnRamp Bioinformatics, Inc., a genomics company providing the premier scientist-focused data analysis platform, and Advaita BioInformatics, a leader in personalized medicine and interpretation of Next-Gen Sequencing data, announced they are partnering to provide a comprehensive research experience from sample to interpretation with a seamless handoff between systems.
OnRamp.Bio’s flagship product, ROSALIND™, enables researchers, drug developers and bench scientists to analyze raw genomics data by providing a transformative experience through point-and-click experiment set up, interactive data visualization and interpretation. This new approach increases productivity by freeing up time for the bioinformatician to focus on more challenging workloads, while making bioinformatic analysis more accessible for the scientist to do more discovery with their data.
Topics: Big Data Personalized Medicine Cancer BioInformatics Biology Rosalind advaita
2 min read
Pipelines & Data Integration
By Jeremy Davis-Turak on Feb 15, 2017 10:36:00 AM
Challenges and opportunities in the research setting.
The emergence and mass utilization of high-throughput (HT) technologies, including sequencing technologies (genomics) and mass spectrometry (proteomics, metabolomics, lipids), has allowed geneticists, biologists, and biostatisticians to bridge the gap between genotype and phenotype on a massive scale. These new technologies have brought rapid advances in our understanding of cell biology, evolutionary history, microbial environments, and are increasingly providing new insights and applications towards clinical care and personalized medicine.
Topics: Big Data Pipelines BioInformatics Genomic Data Management
2 min read
MICROBIOMES & OUR HEALTH
By Jeremy Davis-Turak on Dec 19, 2016 1:30:00 PM
Could health outcomes be determined by the little guys who call us home?
The San Diego MicroBiome Meetup, hosted by Clotide Telling of Illumina, Inc., kicked off with a presentation by OnRamp's Scientific Advisor, Rob Edwards. Rob's story of the discovery of the crAssphage is a truly amazing example of Metagenomics and the MicroBiome, and is testament to the power of the sequencing revolution.
Topics: microbiome crAssphage San Diego State University
3 min read
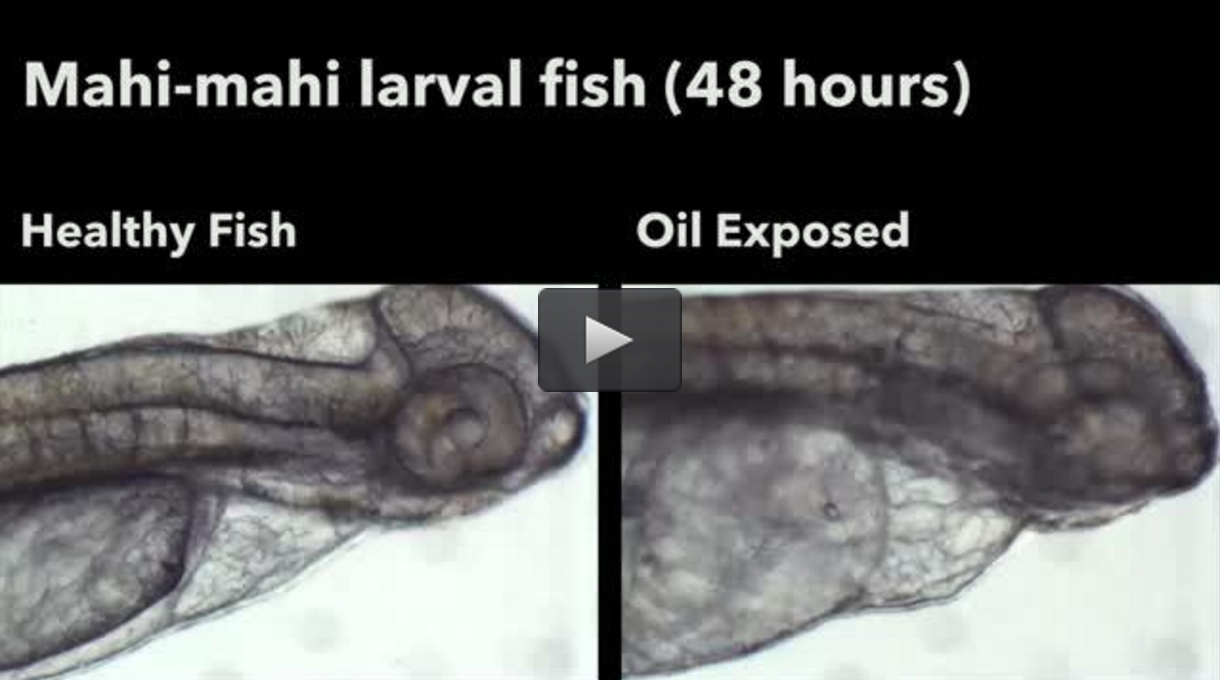
Horizon Oil Spill Effects on Mahi-Mahi | Insights powered by Rosalind
By Jeremy Davis-Turak on Jul 20, 2016 9:38:00 AM
The threat to fish embryos and larvae development.
MIAMI-A research team led by scientists at the University of California, Riverside and the University of Miami (UM) Rosenstiel School of Marine and Atmospheric Science have found that ultraviolet light is changing the structure of the Deepwater Horizon (DWH) oil components into something more toxic, further threatening numerous commercially and ecologically important fishes.
Topics: BioInformatics Genomic Data Management
2 min read
OnRamp Featured in Frontline Genomics Magazine
By Jeremy Davis-Turak on Jan 20, 2016 11:44:00 AM